"The Hidden Flexibility of the Genetic Code and its Untapped Potential"
 Dr. Veronika Kivenson
Dr. Veronika Kivenson
Bio:
Veronika Kivenson is a microbiologist studying genetic code plasticity in bacteria and archaea. Her research examines these processes in diverse settings, ranging from deep-sea sediments to the human microbiome. She earned her Ph.D. from UC Santa Barbara with support from an NSF Graduate Research Fellowship, followed by a Simons Foundation Postdoctoral Fellowship at Oregon State University. Most recently, as a postdoc at UC Berkeley, she characterized the first alternative genetic code in Archaea; this work was published in Science. Her future research focuses on uncovering new genetic codes, the evolutionary drivers of code flexibility and their applications.
Abstract:
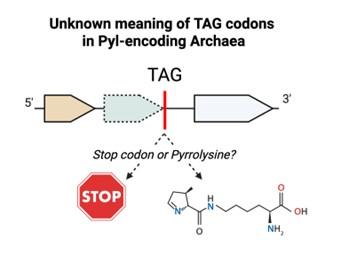
Microbes play a central role in global ecosystems and human health. Their evolutionary success relies on various molecular strategies, including genetic code flexibility. This suite of processes, which includes recoding, frameshifting and codon reassignment, allows them to dynamically interpret genetic information and create novel functions. These processes, however, are largely overlooked: Standard genomic tools are built on the assumption of a largely static, universal code. My recent work demonstrates that alternative genetic codes can evolve rapidly, suggesting that the extent of genetic code flexibility has been significantly underestimated. In the course of that study, I also uncovered numerous other cases of genetic code flexibility and developed the computational framework to find these occurrences systematically. My research program will focus on defining the full extent of genetic code flexibility, characterizing its evolutionary drivers and applying this fundamental knowledge to address challenges in climate change, human health and synthetic biology.

 Dr. Brittany Belin
Dr. Brittany Belin Dr. Stefan Katharios-Lanwermeyer
Dr. Stefan Katharios-Lanwermeyer