Fall 2025 Faculty Spotlight
 Dr. Alexandra Long | Long Lab
Dr. Alexandra Long | Long Lab
How long have you been with the department?
 Dr. Alexandra Long | Long Lab
Dr. Alexandra Long | Long LabHow long have you been with the department?
 Sophia Novoa
Sophia NovoaWhat is your major?
I’m currently a biology major.
Minors?
Political science.

Bio:
Brent Simpson trained at the Ohio State University, where he received his Ph.D. working in the laboratory of Natacha Ruiz. During his Ph.D., he studied how Gram-negative bacteria build the outer membrane and published three lead-author papers exploring the mechanism of LPS transport. He also received awards for both excellence in teaching and a distinguished dissertation during his graduate work. Brent then moved to the University of Georgia to work with Stephen Trent.
He began working on why LPS is essential in most Gram-negative bacteria using a rare model organism that is able to survive with complete absence of its version of LPS that is missing O-antigen and called LOS. He performed the first transposon-sequencing in a Gram-negative organism completely missing LPS or LOS. This experiment revealed new details about multiple aspects of peptidoglycan physiology in the Gram-negative pathogen Acinetobacter baumannii.
He has published four papers in mBio, PNAS and Journal of Bacteriology following up on peptidoglycan physiology. In addition, he was awarded an NIH F32 Kirschstein fellowship and the UGA Postdoctoral Research Award for his research contributions. He plans to leverage his data set to learn more about cell physiology of Acinetobacter and other Gram negatives with the hope of developing strategies to overcome multi-drug resistant infections. Brent is passionate about microbial physiology, mentoring and teaching students, and his three cats.
Abstract:
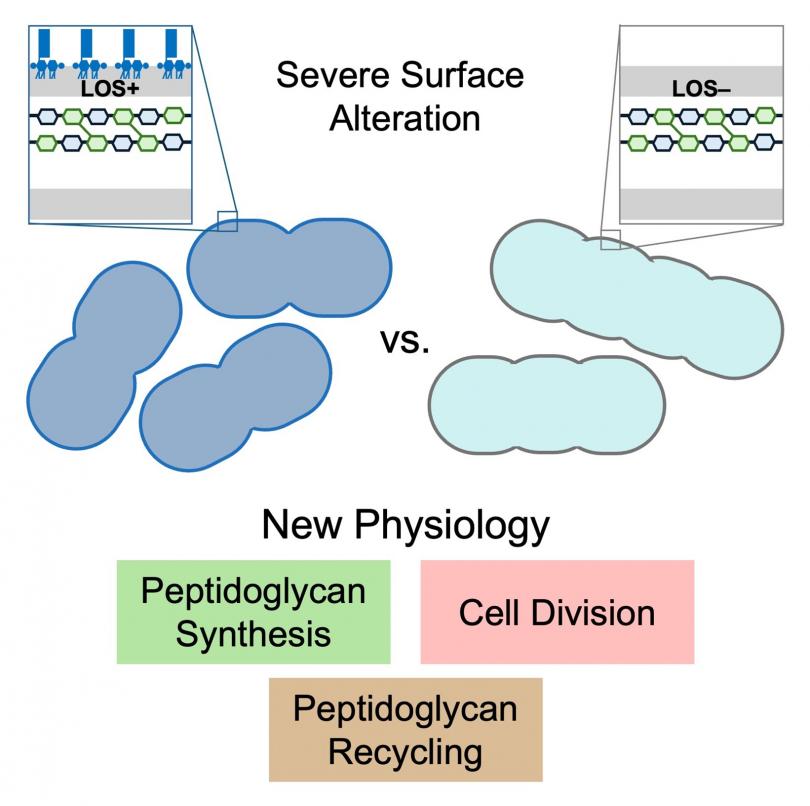
Bacteria have evolved different cell envelope compositions that provide them specific advantages in their natural environments or as human pathogens. Gram-negative bacteria have a three-layered cell envelope with an inner membrane, peptidoglycan, and outer membrane.
The outer membrane has asymmetric organization of lipopolysaccharide (LPS) or lipooligosaccharide (LOS) only on the cell surface. LPS and LOS differ only in the presence (LPS) or absence (LOS) of an O-antigen. LPS/LOS is essential in most Gram negatives serving both as a permeability barrier and providing rigidity to the cell surface. Peptidoglycan is a mesh of long polysaccharide chains crosslinked with short peptides, that also provides rigidity to the cell. The outer membrane and peptidoglycan are critical structures of the cell that if targeted by antimicrobial therapy can cause bacterial clearance.
The high-priority Gram-negative pathogen Acinetobacter baumannii normally produces an LOS-containing outer membrane, but it is remarkably able to survive even with complete loss of LOS. We have used the comparison of A. baumannii with and without LOS as a critical tool to understand how it builds its cell envelope. A transposon sequencing approach in A. baumannii completely missing LOS unveiled new aspects of peptidoglycan synthesis, recycling, and degradation during cell division. Understanding the biogenesis of the outer membrane and peptidoglycan has broad implications in bacterial pathogenesis, antimicrobial resistance and the development of new therapeutic strategies.
 Dr. Veronika Kivenson
Dr. Veronika KivensonBio:
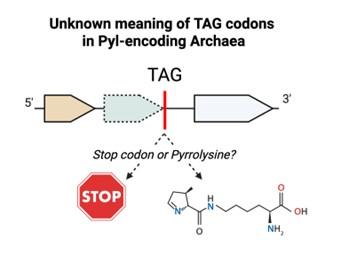
Veronika Kivenson is a microbiologist studying genetic code plasticity in bacteria and archaea. Her research examines these processes in diverse settings, ranging from deep-sea sediments to the human microbiome. She earned her Ph.D. from UC Santa Barbara with support from an NSF Graduate Research Fellowship, followed by a Simons Foundation Postdoctoral Fellowship at Oregon State University. Most recently, as a postdoc at UC Berkeley, she characterized the first alternative genetic code in Archaea; this work was published in Science. Her future research focuses on uncovering new genetic codes, the evolutionary drivers of code flexibility and their applications.
Abstract:
Microbes play a central role in global ecosystems and human health. Their evolutionary success relies on various molecular strategies, including genetic code flexibility. This suite of processes, which includes recoding, frameshifting and codon reassignment, allows them to dynamically interpret genetic information and create novel functions. These processes, however, are largely overlooked: Standard genomic tools are built on the assumption of a largely static, universal code. My recent work demonstrates that alternative genetic codes can evolve rapidly, suggesting that the extent of genetic code flexibility has been significantly underestimated. In the course of that study, I also uncovered numerous other cases of genetic code flexibility and developed the computational framework to find these occurrences systematically. My research program will focus on defining the full extent of genetic code flexibility, characterizing its evolutionary drivers and applying this fundamental knowledge to address challenges in climate change, human health and synthetic biology.
 Dr. Brittany Belin
Dr. Brittany BelinBio:
Dr.Brittany Belin is a graduate of Notre Dame (B.S., biochemistry & philosophy) and the University of California San Francisco (Ph.D., bBiophysics), where she studied the actin cytoskeleton using quantitative microscopy, cell biology and computational analysis. For her postdoc work, she transitioned to studying plant-bacteria symbiosis at Caltech, and she opened her own lab studying lipid biophysics in plant-bacteria symbiosis at the Carnegie Institution for Science in 2020. She is interested in symbiosis and microbial cell biology in all of its forms, and she loves learning and teaching about emerging models for studying these topics.
Abstract:
Life is powered by fixed carbon and nitrogen, and the exchange of these elements is a common basis for bacteria-eukaryote symbioses. The Bradyrhizobia are a globally dominant soil bacteria that form nitrogen-carbon exchange symbioses with peanuts, soybeans and other legumes, and work in our lab and others has shown that these symbioses require cholesterol-like bacterial lipids known as hopanoids.
The mechanisms through which hopanoids participate in symbiosis are not clear, but our preliminary data suggest that hopanoids:
These findings suggest a fundamental connection between bacterial membrane biophysics and successful host-microbe interactions that likely is mediated by diverse, membrane-rigidifying “secondary lipids” throughout the bacterial domain of life.
 Dr. Stefan Katharios-Lanwermeyer
Dr. Stefan Katharios-LanwermeyerBio:
Dr. Stefan Katharios-Lanwermeyer is a postdoctoral researcher at the National Institutes of Health where he supported a PRAT postdoctoral Fellowship. He received his PhD in Microbiology from Dartmouth College where he studied the maintenance of bacterial biofilm under his mentor, Dr. George O’Toole. Currently, Katharios studies bacterial multicellularity in polymicrobial environments under his postdoctoral mentor Dr. Anupama Khare. His goal is to develop a model system to characterize how interspecies interactions result in multicellularity among cohabitating bacteria.
Abstract:
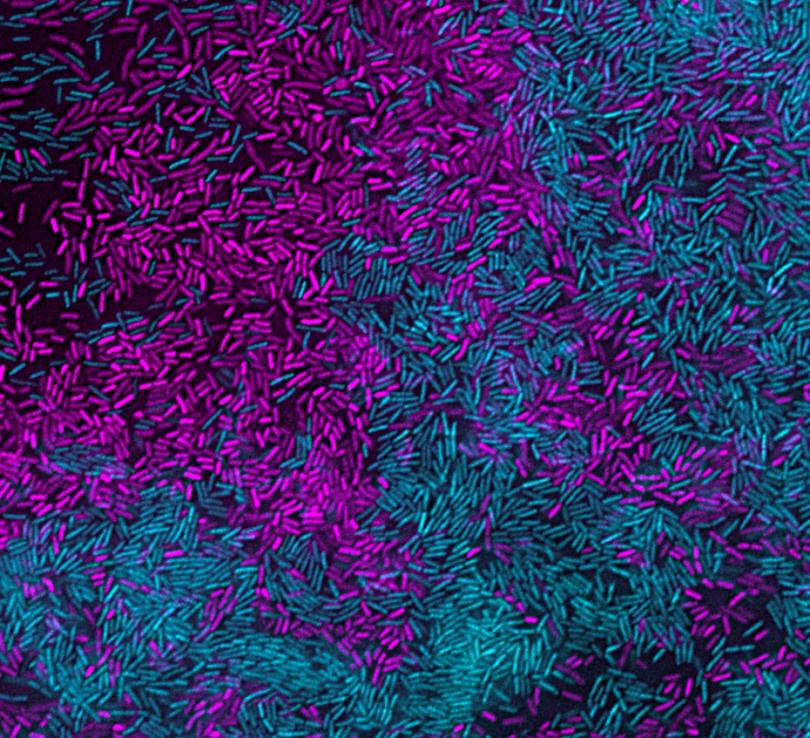
Microorganisms commonly exist in polymicrobial communities, where they can respond to interspecies secreted molecules by altering behaviors and physiology. However, the underlying mechanisms remain underexplored. I have investigated interactions between Stenotrophomonas maltophilia and Pseudomonas aeruginosa, co-infecting bacterial species pathogens found in pneumonia and chronic lung infections, such as in cystic fibrosis. We found that S. maltophilia forms large protective multicellular aggregates upon exposure to P. aeruginosa secreted factors. E
Experimental evolution for lack of aggregation selected for fimbrial mutations, and we found that fimbriae are required on both interacting S. maltophilia cells for aggregation. Untargeted metabolomics and targeted validations revealed that the quorum sensing molecule Pseudomonas quinolone signal (PQS) directly induced S. maltophilia aggregation, and co-localized with the aggregates. Further, in co-culture with P. aeruginosa, wild-type S. maltophilia formed aggregates, resulting in up to 75-fold increased survival from P. aeruginosa competition compared to fimbrial mutants.
Finally, multiple other bacterial species similarly aggregated upon exposure to P. aeruginosa PQS, indicating a more general response. Collectively, our work identifies a novel multispecies interaction where a quorum sensing molecule from a co-infecting pathogen is sensed as a ‘danger’ signal, thereby inducing a protective multicellular response. This work provides a basis to expand to a three-species model system in which I aim to investigate bacterial multicellularity and interspecies interactions as bacteria respond to secreted factors produced by cohabitating microbes.
 Michael Rossi, Ph.D.
Michael Rossi, Ph.D.What is your connection to the Department of Biology?


View Katz's biosketch here.
Abstract:
The Katz lab is focused on the heritability of histone modifications and how this heritability contributes to traits and disease. We demonstrated in C. elegans how histone modifications can be trans-generationally maintained and lead to heritable phenotypes ranging from sterility and longevity to behavior.
These phenotypes demonstrate the histone modifications can serve as heritable information across generations. Based on these data, we have engineered a new mouse model where maternal epigenetic reprograming activity of the H3K4me1/2 demethylase LSD1/KDM1A is compromised.
These maternally compromised mice exhibit inherited phenotypes that manifest postnatally, including perinatal lethality, developmental delay, craniofacial defects and altered behavior. These phenotypes include all of those found in the corresponding Kabuki-syndrome-like human patients, raising the possibility that maternal defects may contribute to phenotypes in LSD1 patients and Kabuki syndrome.
Finally, we made the surprising discovery that loss of LSD1 in adult mice leads to neurodegeneration. Following up on this result, we generated significant data suggesting that LSD1 functions specifically in the pathological tau pathway. Pathological tau is thought to be a critical driver of neurodegeneration in Alzheimer’s disease. Based on our data, we propose that pathological tau contributes to neuronal cell death in Alzheimer’s disease by sequestering LSD1 in the cytoplasm and interfering with the continuous requirement for LSD1 to epigenetically repress transcription associated with alternative cell fates. Thus, it may be possible to target LSD1 therapeutically to block tau-mediated neurodegeneration.
Watch the seminar here!